5 Functions
5.1 Function fundamentals
Q1: Given a name, like "mean", match.fun() lets you find a function. Given a function, can you find its name? Why doesn’t that make sense in R?
A: In R there is no one-to-one mapping between functions and names. A name always points to a single object, but an object may have zero, one or many names.
Let’s look at an example:
function(x) sd(x) / mean(x)
#> function(x) sd(x) / mean(x)
f1 <- function(x) (x - min(x)) / (max(x) - min(x))
f2 <- f1
f3 <- f1While the function in the first line is not bound to a name multiple names (f1, f2 and f3) point to the second function. So, the main point is that the relation between name and object is only clearly defined in one direction.
Besides that, there are obviously ways to search for function names. However, to be sure to find the right one(s), you should not only compare the code (body) but also the arguments (formals) and the creation environment. As formals(), body() and environment() all return NULL for primitive functions, the easiest way to check if two functions are exactly equal is just to use identical().
Q2: It’s possible (although typically not useful) to call an anonymous function. Which of the two approaches below is correct? Why?
function(x) 3()
#> function(x) 3()
(function(x) 3)()
#> [1] 3A: The second approach is correct.
The anonymous function function(x) 3 is surrounded by a pair of parentheses before it is called by (). These extra parentheses separate the function call from the anonymous function’s body. Without them a function with the invalid body 3() is returned, which throws an error when we call it. This is easier to see if we name the function:
f <- function(x) 3()
f
#> function(x) 3()
f()
#> Error in f(): attempt to apply non-functionQ3: A good rule of thumb is that an anonymous function should fit on one line and shouldn’t need to use {}. Review your code. Where could you have used an anonymous function instead of a named function? Where should you have used a named function instead of an anonymous function?
A: The use of anonymous functions allows concise and elegant code in certain situations. However, they miss a descriptive name and when re-reading the code, it can take a while to figure out what they do. That’s why it’s helpful to give long and complex functions a descriptive name. It may be worthwhile to take a look at your own projects or other people’s code to reflect on this part of your coding style.
Q4: What function allows you to tell if an object is a function? What function allows you to tell if a function is a primitive function?
A: Use is.function() to test if an object is a function. Consider using is.primitive() to test specifically for primitive functions.
Q5: This code makes a list of all functions in the {base} package.
Use it to answer the following questions:
Which base function has the most arguments?
How many base functions have no arguments? What’s special about those functions?
How could you adapt the code to find all primitive functions?
A: Let’s look at each sub-question separately:
-
To find the function with the most arguments, we first compute the length of
formals().Then we sort
n_argsin decreasing order and look at its first entries. -
We can further use
n_argsto find the number of functions with no arguments:sum(n_args == 0) #> [1] 248However, this over counts because
formals()returnsNULLfor primitive functions, andlength(NULL)is 0. To fix this, we can first remove the primitive functions:n_args2 <- funs %>% discard(is.primitive) %>% map(formals) %>% map_int(length) sum(n_args2 == 0) #> [1] 47Indeed, most of the functions with no arguments are actually primitive functions.
-
To find all primitive functions, we can change the predicate in
Filter()fromis.function()tois.primitive():
Q6: What are the three important components of a function?
A: These components are the function’s body(), formals() and environment(). However, as mentioned in Advanced R:
There is one exception to the rule that functions have three components. Primitive functions, like
sum(), call C code directly with.Primitive()and contain no R code. Therefore, theirformals(),body(), andenvironment()are allNULL.
Q7: When does printing a function not show what environment it was created in?
A: Primitive functions and functions created in the global environment do not print their environment.
5.2 Lexical scoping
Q1: What does the following code return? Why? Describe how each of the three c’s is interpreted.
c <- 10
c(c = c)A: This code returns a named numeric vector of length one — with one element of the value 10 and the name "c". The first c represents the c() function, the second c is interpreted as a (quoted) name and the third c as a value.
Q2: What are the four principles that govern how R looks for values?
A: R’s lexical scoping rules are based on these four principles:
Q3: What does the following function return? Make a prediction before running the code yourself.
f <- function(x) {
f <- function(x) {
f <- function() {
x ^ 2
}
f() + 1
}
f(x) * 2
}
f(10)A: Within this nested function two more functions also named f are defined and called. Because the functions are each executed in their own environment R will look up and use the functions defined last in these environments. The innermost f() is called last, though it is the first function to return a value. Therefore, the order of the calculation passes “from the inside to the outside” and the function returns ((10 ^ 2) + 1) * 2, i.e. 202.
5.3 Lazy evaluation
Q1: What important property of && makes x_ok() work?
x_ok <- function(x) {
!is.null(x) && length(x) == 1 && x > 0
}
x_ok(NULL)
#> [1] FALSE
x_ok(1)
#> [1] TRUE
x_ok(1:3)
#> [1] FALSEWhat is different with this code? Why is this behaviour undesirable here?
x_ok <- function(x) {
!is.null(x) & length(x) == 1 & x > 0
}
x_ok(NULL)
#> logical(0)
x_ok(1)
#> [1] TRUE
x_ok(1:3)
#> [1] FALSE FALSE FALSEA: In summary: && short-circuits which means that if the left-hand side is FALSE it doesn’t evaluate the right-hand side (because it doesn’t matter). Similarly, if the left-hand side of || is TRUE it doesn’t evaluate the right-hand side.
We expect x_ok() to validate its input via certain criteria: it must not be NULL, have length 1 and be greater than 0. Meaningful outcomes for this assertion will be TRUE, FALSE or NA. The desired behaviour is reached by combining the assertions through && instead of &.
&& does not perform elementwise comparisons; instead it uses the first element of each value only. It also uses lazy evaluation, in the sense that evaluation “proceeds only until the result is determined” (from ?Logic). This means that the RHS of && won’t be evaluated if the LHS already determines the outcome of the comparison (e.g. evaluate to FALSE). This behaviour is also known as “short-circuiting.”
For some situations (x = 1) both operators will lead to the same result. But this is not always the case. For x = NULL, the &&-operator will stop after the !is.null statement and return the result. The following conditions won’t even be evaluated! (If the other conditions are also evaluated (by the use of &), the outcome would change. NULL > 0 returns logical(0), which is not helpful in this case.)
We can also see the difference in behaviour, when we set x = 1:3. The &&-operator returns the result from length(x) == 1, which is FALSE. Using & as the logical operator leads to the (vectorised) x > 0 condition being evaluated and also returned.
Q2: What does this function return? Why? Which principle does it illustrate?
f2 <- function(x = z) {
z <- 100
x
}
f2()A: The function returns 100. The default argument (x = z) gets lazily evaluated within the function environment when x gets accessed. At this time z has already been bound to the value 100. The illustrated principle here is lazy evaluation.
Q3: What does this function return? Why? Which principle does it illustrate?
y <- 10
f1 <- function(x = {y <- 1; 2}, y = 0) {
c(x, y)
}
f1()
yA: The function returns c(2, 1) which is due to name masking. When x is accessed within c(), the promise x = {y <- 1; 2} is evaluated inside f1()’s environment. y gets bound to the value 1 and the return value of {() (2) gets assigned to x. When y gets accessed next within c(), it has already the value 1 and R doesn’t need to look it up any further. Therefore, the promise y = 0 won’t be evaluated. Also, as y is assigned within f1()’s environment, the value of the global variable y is left untouched.
Q4: In hist(), the default value of xlim is range(breaks), the default value for breaks is "Sturges", and
range("Sturges")
#> [1] "Sturges" "Sturges"Explain how hist() works to get a correct xlim value.
A: The xlim argument of hist() defines the range of the histogram’s x-axis. In order to provide a valid axis xlim must contain a numeric vector of exactly two unique values. Consequently, for the default xlim = range(breaks)), breaks must evaluate to a vector with at least two unique values.
During execution hist() overwrites the breaks argument. The breaks argument is quite flexible and allows the users to provide the breakpoints directly or compute them in several ways. Therefore, the specific behaviour depends highly on the input. But hist ensures that breaks evaluates to a numeric vector containing at least two unique elements before xlim is computed.
Q5: Explain why this function works. Why is it confusing?
show_time <- function(x = stop("Error!")) {
stop <- function(...) Sys.time()
print(x)
}
show_time()
#> [1] "2021-05-02 09:44:05 UTC"A: Before show_time() accesses x (default stop("Error")), the stop() function is masked by function(...) Sys.time(). As default arguments are evaluated in the function environment, print(x) will be evaluated as print(Sys.time()).
This function is confusing because its behaviour changes when x’s value is supplied directly. Now the value from the calling environment will be used and the overwriting of stop() won’t affect x anymore.
show_time(x = stop("Error!"))
#> Error in print(x): Error!Q6: How many arguments are required when calling library()?
A: library() doesn’t require any arguments. When called without arguments library() invisibly returns a list of class libraryIQR, which contains a results matrix with one row and three columns per installed package. These columns contain entries for the name of the package (“Package”), the path to the package (“LibPath”) and the title of the package (“Title”). library() also has its own print method (print.libraryIQR()), which displays this information conveniently in its own window.
This behaviour is also documented under the details section of library()’s help page (?library):
If library is called with no package or help argument, it lists all available packages in the libraries specified by lib.loc, and returns the corresponding information in an object of class “libraryIQR.” (The structure of this class may change in future versions.) Use .packages(all = TRUE) to obtain just the names of all available packages, and installed.packages() for even more information.
Because the package and help argument from library() do not show a default value, it’s easy to overlook the possibility to call library() without these arguments. (Instead of providing NULLs as default values library() uses missing() to check if these arguments were provided.)
str(formals(library))
#> Dotted pair list of 13
#> $ package : symbol
#> $ help : symbol
#> $ pos : num 2
#> $ lib.loc : NULL
#> $ character.only : logi FALSE
#> $ logical.return : logi FALSE
#> $ warn.conflicts : symbol
#> $ quietly : logi FALSE
#> $ verbose : language getOption("verbose")
#> $ mask.ok : symbol
#> $ exclude : symbol
#> $ include.only : symbol
#> $ attach.required: language missing(include.only)
5.4 ... (dot-dot-dot)
Q1: Explain the following results:
sum(1, 2, 3)
#> [1] 6
mean(1, 2, 3)
#> [1] 1
sum(1, 2, 3, na.omit = TRUE)
#> [1] 7
mean(1, 2, 3, na.omit = TRUE)
#> [1] 1A: Let’s inspect the arguments and their order for both functions. For sum() these are ... and na.rm:
str(sum)
#> function (..., na.rm = FALSE)For the ... argument sum() expects numeric, complex, or logical vector input (see ?sum). Unfortunately, when ... is used, misspelled arguments (!) like na.omit won’t raise an error (in case of no further input checks). So instead, na.omit is treated as a logical and becomes part of the ... argument. It will be coerced to 1 and be part of the sum. All other arguments are left unchanged. Therefore sum(1, 2, 3) returns 6 and sum(1, 2, 3, na.omit = TRUE) returns 7.
In contrast, the generic function mean() expects x, trim, na.rm and ... for its default method.
str(mean.default)
#> function (x, trim = 0, na.rm = FALSE, ...)As na.omit is not one of mean()’s named arguments (x; and no candidate for partial matching), na.omit again becomes part of the ... argument. However, in contrast to sum() the elements of ... are not “part” of the mean. The other supplied arguments are matched by their order, i.e. x = 1, trim = 2 and na.rm = 3. As x is of length 1 and not NA, the settings of trim and na.rm do not affect the calculation of the mean. Both calls (mean(1, 2, 3) and mean(1, 2, 3, na.omit = TRUE)) return 1.
Q2: Explain how to find the documentation for the named arguments in the following function call:
plot(1:10, col = "red", pch = 20, xlab = "x", col.lab = "blue")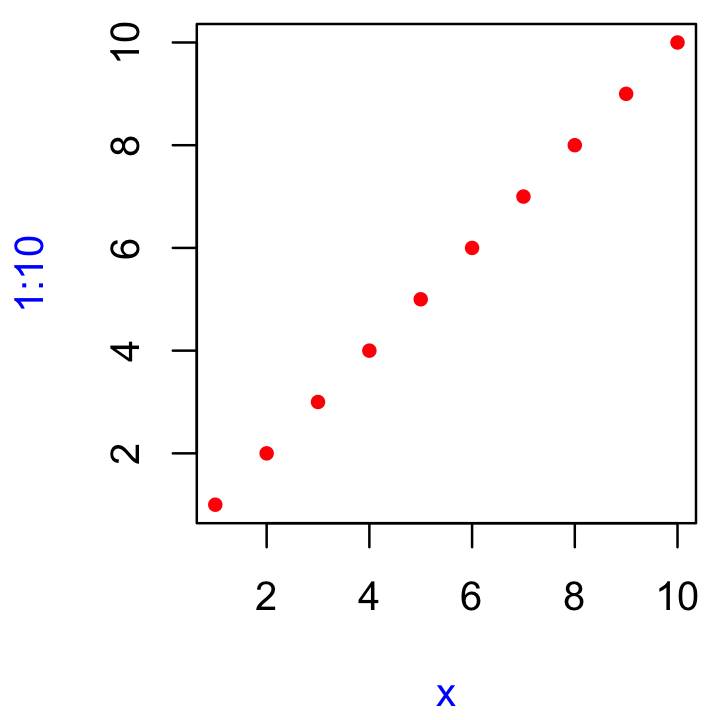
A: First we type ?plot in the console and check the “Usage” section which contains:
plot(x, y, ...)The arguments we want to learn more about (col, pch, xlab, col.lab) are part of the ... argument. There we can find information for the xlab argument and a recommendation to visit ?par for the other arguments. Under ?par we type “col” into the search bar, which leads us to the section “Color Specification.” We also search for the pch argument, which leads to the recommendation to check ?points. Finally, col.lab is also directly documented within ?par.
Q3: Why does plot(1:10, col = "red") only colour the points, not the axes or labels? Read the source code of plot.default() to find out.
A: To learn about the internals of plot.default() we add browser() to the first line of the code and interactively run plot(1:10, col = "red"). This way we can see how the plot is built and learn where the axes are added.
This leads us to the function call
localTitle(main = main, sub = sub, xlab = xlab, ylab = ylab, ...)The localTitle() function was defined in the first lines of plot.default() as:
localTitle <- function(..., col, bg, pch, cex, lty, lwd) title(...)The call to localTitle() passes the col parameter as part of the ... argument to title(). ?title tells us that the title() function specifies four parts of the plot: Main (title of the plot), sub (sub-title of the plot) and both axis labels. Therefore, it would introduce ambiguity inside title() to use col directly. Instead, one has the option to supply col via the ... argument, via col.lab or as part of xlab in the form xlab = list(c("index"), col = "red") (similar for ylab).
5.5 Exiting a function
Q1: What does load() return? Why don’t you normally see these values?
A: load() loads objects saved to disk in .Rdata files by save(). When run successfully, load() invisibly returns a character vector containing the names of the newly loaded objects. To print these names to the console, one can set the argument verbose to TRUE or surround the call in parentheses to trigger R’s auto-printing mechanism.
Q2: What does write.table() return? What would be more useful?
A: write.table() writes an object, usually a data frame or a matrix, to disk. The function invisibly returns NULL. It would be more useful if write.table() would (invisibly) return the input data, x. This would allow to save intermediate results and directly take on further processing steps without breaking the flow of the code (i.e. breaking it into different lines). One package which uses this pattern is the {readr} package,12 which is part of the tidyverse-ecosystem.
Q3: How does the chdir parameter of source() compare to with_dir()? Why might you prefer one to the other?
A: The with_dir() approach was given in Advanced R as:
with_dir() takes a path for a working directory (dir) as its first argument. This is the directory where the provided code (code) should be executed. Therefore, the current working directory is changed in with_dir() via setwd(). Then, on.exit() ensures that the modification of the working directory is reset to the initial value when the function exits. By passing the path explicitly, the user has full control over the directory to execute the code in.
In source() the code is passed via the file argument (a path to a file). The chdir argument specifies if the working directory should be changed to the directory containing the file. The default for chdir is FALSE, so you don’t have to provide a value. However, as you can only provide TRUE or FALSE, you are also less flexible in choosing the working directory for the code execution.
Q4: Write a function that opens a graphics device, runs the supplied code, and closes the graphics device (always, regardless of whether or not the plotting code works).
A: To control the graphics device we use pdf() and dev.off(). To ensure a clean termination on.exit() is used.
Q5: We can use on.exit() to implement a simple version of capture.output().
capture.output2 <- function(code) {
temp <- tempfile()
on.exit(file.remove(temp), add = TRUE, after = TRUE)
sink(temp)
on.exit(sink(), add = TRUE, after = TRUE)
force(code)
readLines(temp)
}
capture.output2(cat("a", "b", "c", sep = "\n"))
#> [1] "a" "b" "c"Compare capture.output() to capture.output2(). How do the functions differ? What features have I removed to make the key ideas easier to see? How have I rewritten the key ideas to be easier to understand?
A: Using body(capture.output) we inspect the source code of the original capture.output() function: The implementation for capture.output() is quite a bit longer (39 lines vs. 7 lines).
In capture_output2() the code is simply forced, and the output is caught via sink() in a temporary file. An additional feature of capture_output() is that one can also capture messages by setting type = "message". As this is internally forwarded to sink(), this behaviour (and also sink()’s split argument) could be easily introduced within capture_output2() as well.
The main difference is that capture.output() calls print, i.e. compare the output of these two calls:
capture.output({1})
#> [1] "[1] 1"
capture.output2({1})
#> character(0)5.6 Function forms
Q1: Rewrite the following code snippets into prefix form:
1 + 2 + 3
1 + (2 + 3)
if (length(x) <= 5) x[[5]] else x[[n]]A: Let’s rewrite the expressions to match the exact syntax from the code above. Because prefix functions already define the execution order, we may omit the parentheses in the second expression.
`+`(`+`(1, 2), 3)
`+`(1, `(`(`+`(2, 3)))
`+`(1, `+`(2, 3))
`if`(`<=`(length(x), 5), `[[`(x, 5), `[[`(x, n))Q2: Clarify the following list of odd function calls:
x <- sample(replace = TRUE, 20, x = c(1:10, NA))
y <- runif(min = 0, max = 1, 20)
cor(m = "k", y = y, u = "p", x = x)A: None of these functions provides a ... argument. Therefore, the function arguments are first matched exactly, then via partial matching and finally by position. This leads us to the following explicit function calls:
x <- sample(c(1:10, NA), size = 20, replace = TRUE)
y <- runif(20, min = 0, max = 1)
cor(x, y, use = "pairwise.complete.obs", method = "kendall")Q3: Explain why the following code fails:
A: First, let’s define x and recall the definition of modify() from Advanced R:
x <- 1:3
`modify<-` <- function(x, position, value) {
x[position] <- value
x
}R internally transforms the code, and the transformed code reproduces the error above:
get("x") <- `modify<-`(get("x"), 1, 10)
#> Error in get("x") <- `modify<-`(get("x"), 1, 10) :
#> target of assignment expands to non-language objectThe error occurs during the assignment because no corresponding replacement function, i.e. get<-, exists for get(). To confirm this, we reproduce the error via the following simplified example.
get("x") <- 2
#> Error in get("x") <- 2 :
#> target of assignment expands to non-language objectQ4: Create a replacement function that modifies a random location in a vector.
A: Let’s define random<- like this:
Q5: Write your own version of + that pastes its inputs together if they are character vectors but behaves as usual otherwise. In other words, make this code work:
1 + 2
#> [1] 3
"a" + "b"
#> [1] "ab"A: To achieve this behaviour, we need to override the + operator. We need to take care to not use the + operator itself inside of the function definition, as this would lead to an undesired infinite recursion. We also add b = 0L as a default value to keep the behaviour of + as a unary operator, i.e. to keep + 1 working and not throwing an error.
`+` <- function(a, b = 0L) {
if (is.character(a) && is.character(b)) {
paste0(a, b)
} else {
base::`+`(a, b)
}
}
# Test
+ 1
#> [1] 1
1 + 2
#> [1] 3
"a" + "b"
#> [1] "ab"
# Return back to the original `+` operator
rm(`+`)Q6: Create a list of all the replacement functions found in the {base} package. Which ones are primitive functions? (Hint use apropos())
A: The hint suggests to look for functions with a specific naming pattern: Replacement functions conventionally end on “<-.” We can search for these objects by supplying the regular expression "<-$" to apropos(). apropos() also allows to return the position on the search path (search()) for each of its matches via setting where = TRUE. Finally, we can set mode = function to narrow down our search to relevant objects only. This gives us the following statement to begin with:
repls <- apropos("<-", where = TRUE, mode = "function")
head(repls, 30)
#> 10 10 10
#> ".rowNamesDF<-" "[[<-" "[[<-.data.frame"
#> 10 10 10
#> "[[<-.factor" "[[<-.numeric_version" "[[<-.POSIXlt"
#> 10 10 10
#> "[<-" "[<-.data.frame" "[<-.Date"
#> 10 10 10
#> "[<-.factor" "[<-.numeric_version" "[<-.POSIXct"
#> 10 10 10
#> "[<-.POSIXlt" "@<-" "<-"
#> 10 10 10
#> "<<-" "$<-" "$<-.data.frame"
#> 8 10 10
#> "as<-" "attr<-" "attributes<-"
#> 8 10 10
#> "body<-" "body<-" "class<-"
#> 8 10 10
#> "coerce<-" "colnames<-" "comment<-"
#> 3 10 10
#> "contrasts<-" "diag<-" "dim<-"To restrict repl to names of replacement functions from the {base} package, we select only matches containing the relevant position on the search path.
repls_base <- repls[names(repls) == length(search())]
repls_base
#> 10 10 10
#> ".rowNamesDF<-" "[[<-" "[[<-.data.frame"
#> 10 10 10
#> "[[<-.factor" "[[<-.numeric_version" "[[<-.POSIXlt"
#> 10 10 10
#> "[<-" "[<-.data.frame" "[<-.Date"
#> 10 10 10
#> "[<-.factor" "[<-.numeric_version" "[<-.POSIXct"
#> 10 10 10
#> "[<-.POSIXlt" "@<-" "<-"
#> 10 10 10
#> "<<-" "$<-" "$<-.data.frame"
#> 10 10 10
#> "attr<-" "attributes<-" "body<-"
#> 10 10 10
#> "class<-" "colnames<-" "comment<-"
#> 10 10 10
#> "diag<-" "dim<-" "dimnames<-"
#> 10 10 10
#> "dimnames<-.data.frame" "Encoding<-" "environment<-"
#> 10 10 10
#> "formals<-" "is.na<-" "is.na<-.default"
#> 10 10 10
#> "is.na<-.factor" "is.na<-.numeric_version" "length<-"
#> 10 10 10
#> "length<-.Date" "length<-.difftime" "length<-.factor"
#> 10 10 10
#> "length<-.POSIXct" "length<-.POSIXlt" "levels<-"
#> 10 10 10
#> "levels<-.factor" "mode<-" "mostattributes<-"
#> 10 10 10
#> "names<-" "names<-.POSIXlt" "oldClass<-"
#> 10 10 10
#> "parent.env<-" "regmatches<-" "row.names<-"
#> 10 10 10
#> "row.names<-.data.frame" "row.names<-.default" "rownames<-"
#> 10 10 10
#> "split<-" "split<-.data.frame" "split<-.default"
#> 10 10 10
#> "storage.mode<-" "substr<-" "substring<-"
#> 10 10
#> "units<-" "units<-.difftime"To find out which of these functions are primitives, we first search for these functions via mget() and then subset the result using Filter() and is.primitive().
repls_base_prim <- mget(repls_base, envir = baseenv()) %>%
Filter(is.primitive, .) %>%
names()
repls_base_prim
#> [1] "[[<-" "[<-" "@<-" "<-"
#> [5] "<<-" "$<-" "attr<-" "attributes<-"
#> [9] "class<-" "dim<-" "dimnames<-" "environment<-"
#> [13] "length<-" "levels<-" "names<-" "oldClass<-"
#> [17] "storage.mode<-"Overall the {base} package contains 62 replacement functions of which 17 are primitive functions.
Q7: What are valid names for user-created infix functions?
A: Let’s cite from the section on function forms from Advanced R:
… names of infix functions are more flexible than regular R functions: they can contain any sequence of characters except “%.”
Q8: Create an infix xor() operator.
A: We could create an infix %xor% like this:
`%xor%` <- function(a, b) {
xor(a, b)
}
TRUE %xor% TRUE
#> [1] FALSE
FALSE %xor% TRUE
#> [1] TRUEQ9: Create infix versions of the set functions intersect(), union(), and setdiff(). You might call them %n%, %u%, and %/% to match conventions from mathematics.
A: These infix operators could be defined in the following way. (%/% is chosen instead of %\%, because \ serves as an escape character.)